It's day 25 of the 30 Day Map Challenge and today's theme is "Hexagons".
Challenge Classic: Use hexagonal binning (hexbins) or a hexagonal grid system to visualize your data. Celebrate this beautiful and efficient tessellation!
Given Icon Map's logo consists of hexagons, you can see how important we think they are! Today seems like a good opportunity to explain why I think they are a great tool for presenting large scale geospatial datasets in Power BI.
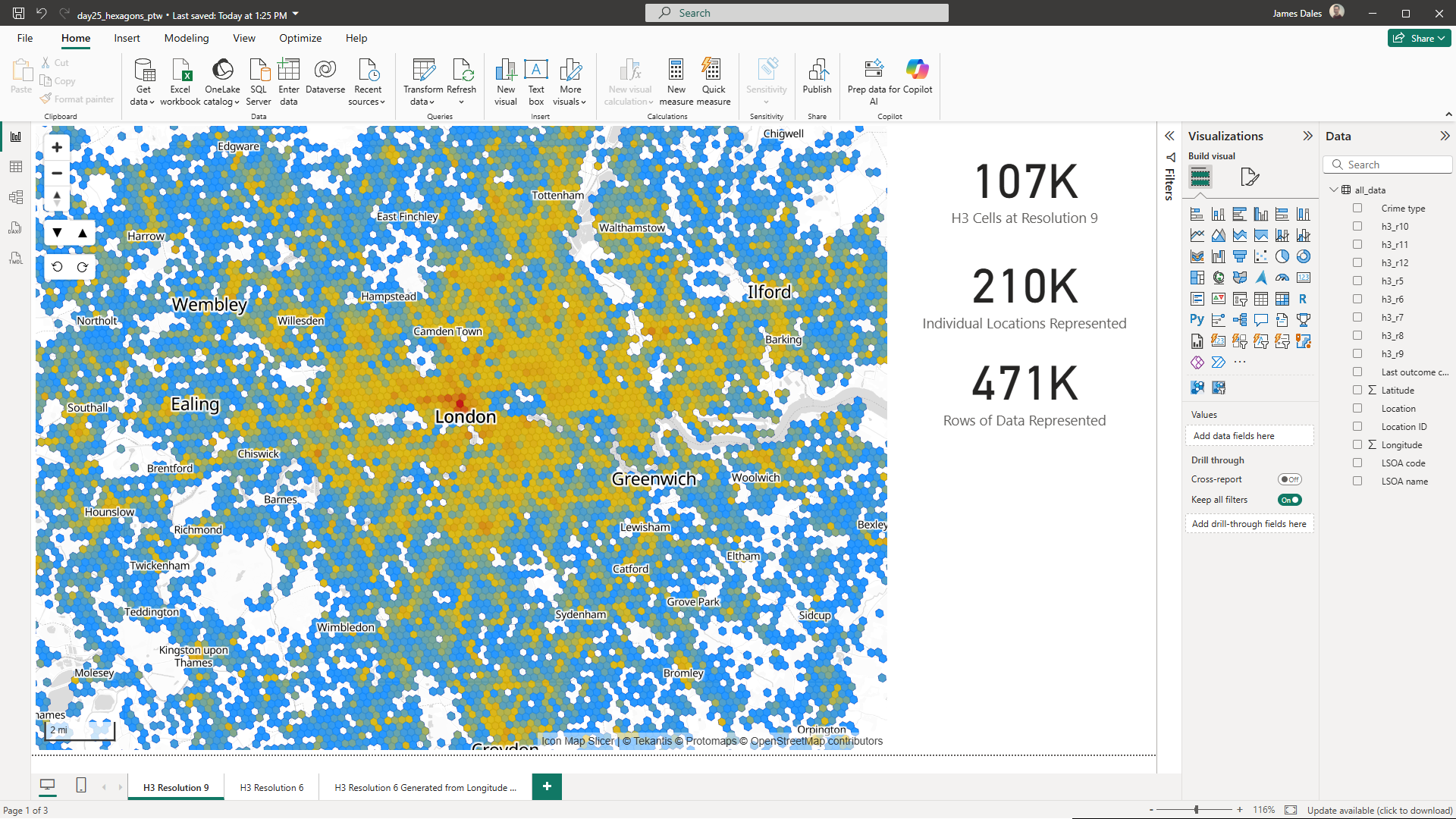
For today's challenge I've downloaded a month's worth of crime data from data.police.uk for all Police Forces. This is around 470,000 rows of data, each row representing a crime.
Most Power BI visuals are limited to 30,000 rows of data, and taking our crime data, even if you pre-aggregate the data, to only send unique locations to the map, it's still over 210,000 locations.
To highlight the issue, here's the Azure Maps visual in Power BI displaying 30,000 locations from our 210,000 location dataset. It only provides a small subset of the full data:
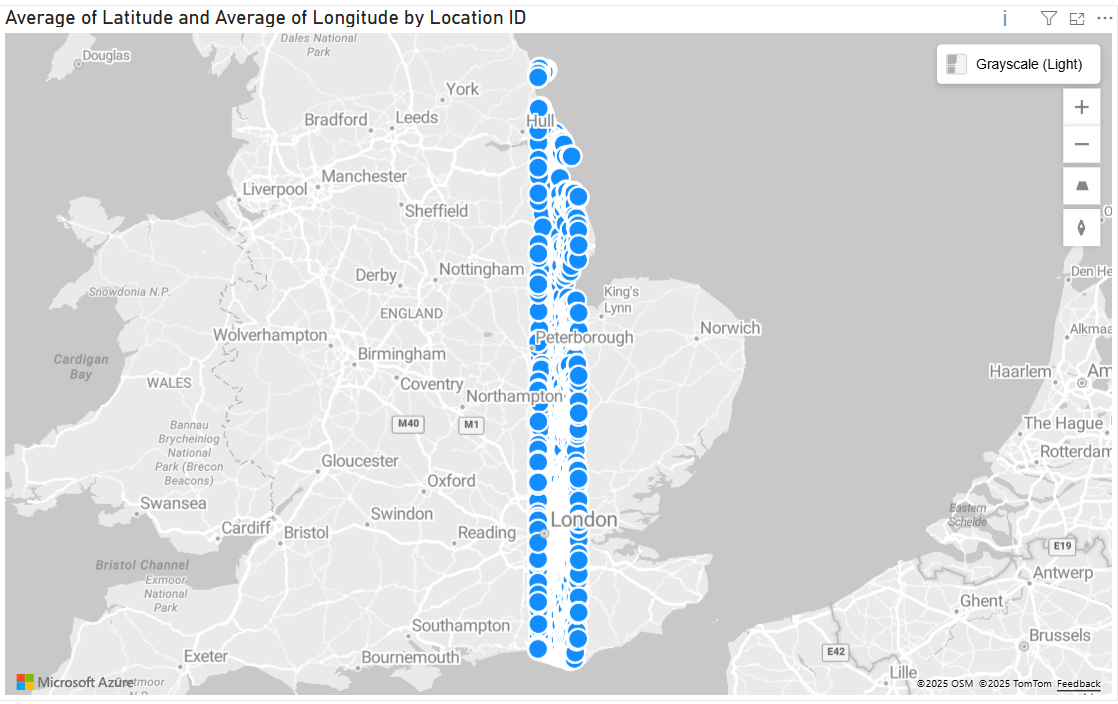
Whilst Icon Map Slicer can display nearly 1/2 million rows, displaying 210,000 locations as circles isn't going to result in an effective visualization:
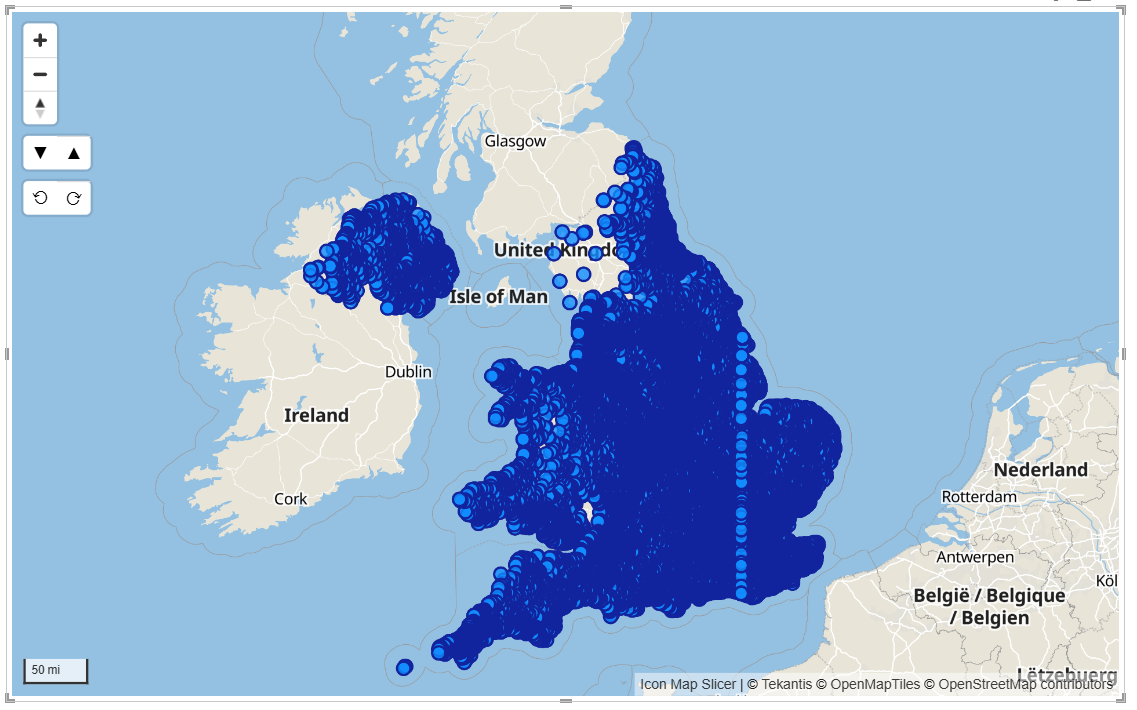
This is where the power of binning our data into hexagons comes into play. Icon Map Slicer has built in support for the H3 hexagon spatial grid. Switching the circle layer to H3 means that Icon Map Slicer can use our longitude and latitude locations, and bin them into H3 hexagon cells, resulting in a much more effective tool for analysis.
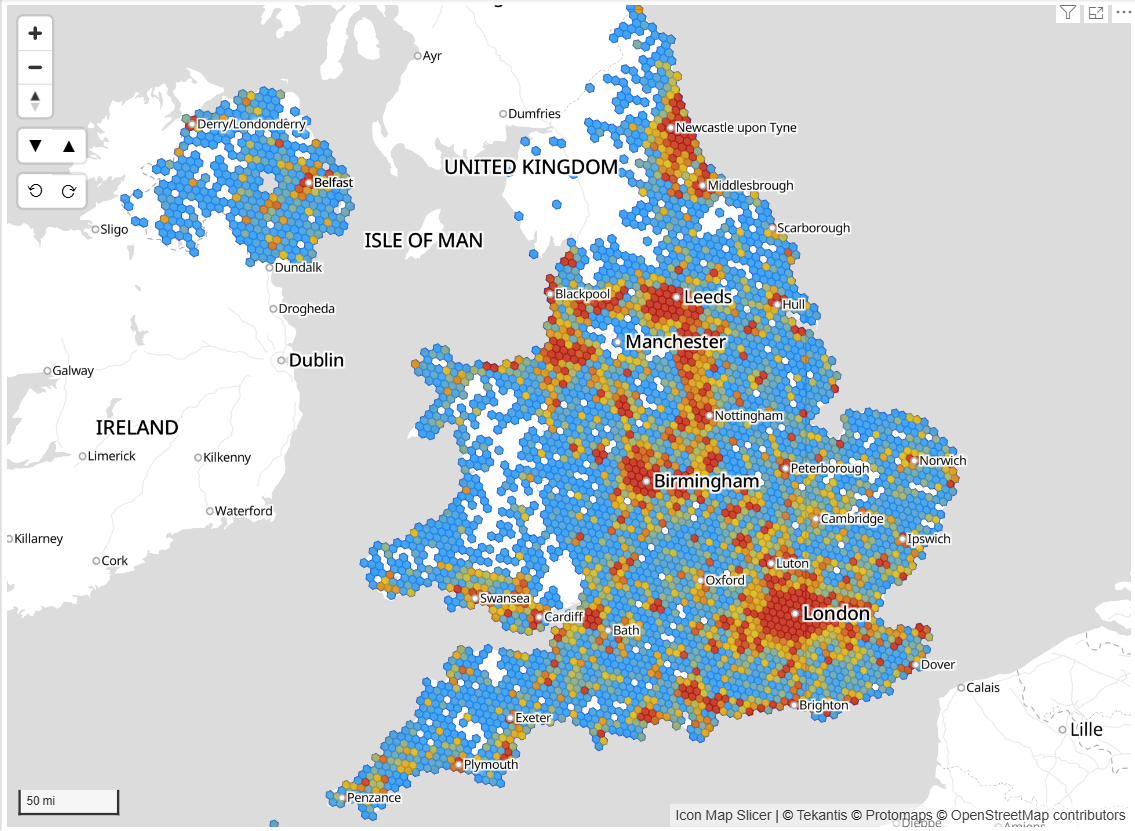
Whilst this works well, it takes Power BI a few seconds to send the data to the visual in 30,000 row segments:
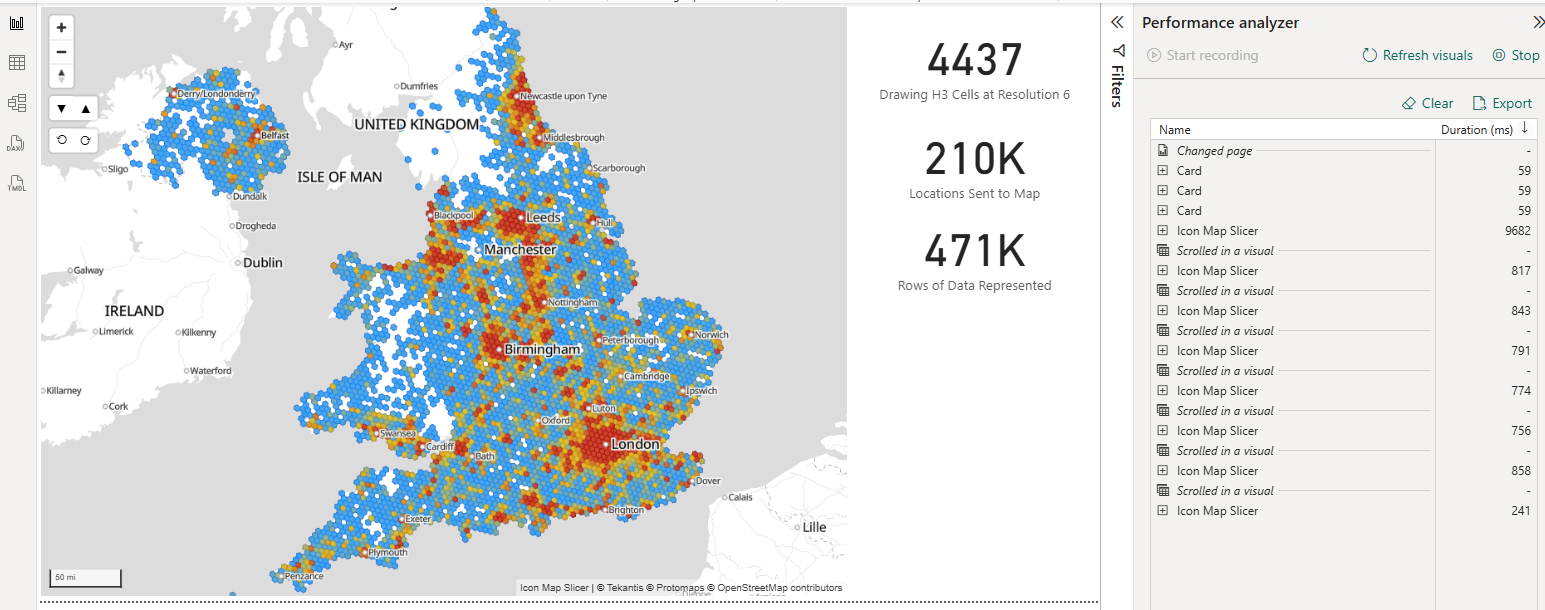
If we're able to pre-aggregate our data into the H3 bins before sending it to Icon Map Pro, then we can drastically increase the speed as we only then need send 1 row per hexagon. For example, our 471,000 rows of crime data can be represented by 4,437 rows - one for each hexagon.
Icon Map Slicer understands the H3 cell IDs representing each hexagon natively - so you don't need to send the GeoJSON to the visual, just the hex or 64 bit integer representing each specific cell. So just 2 columns are required to draw the hexagon map:
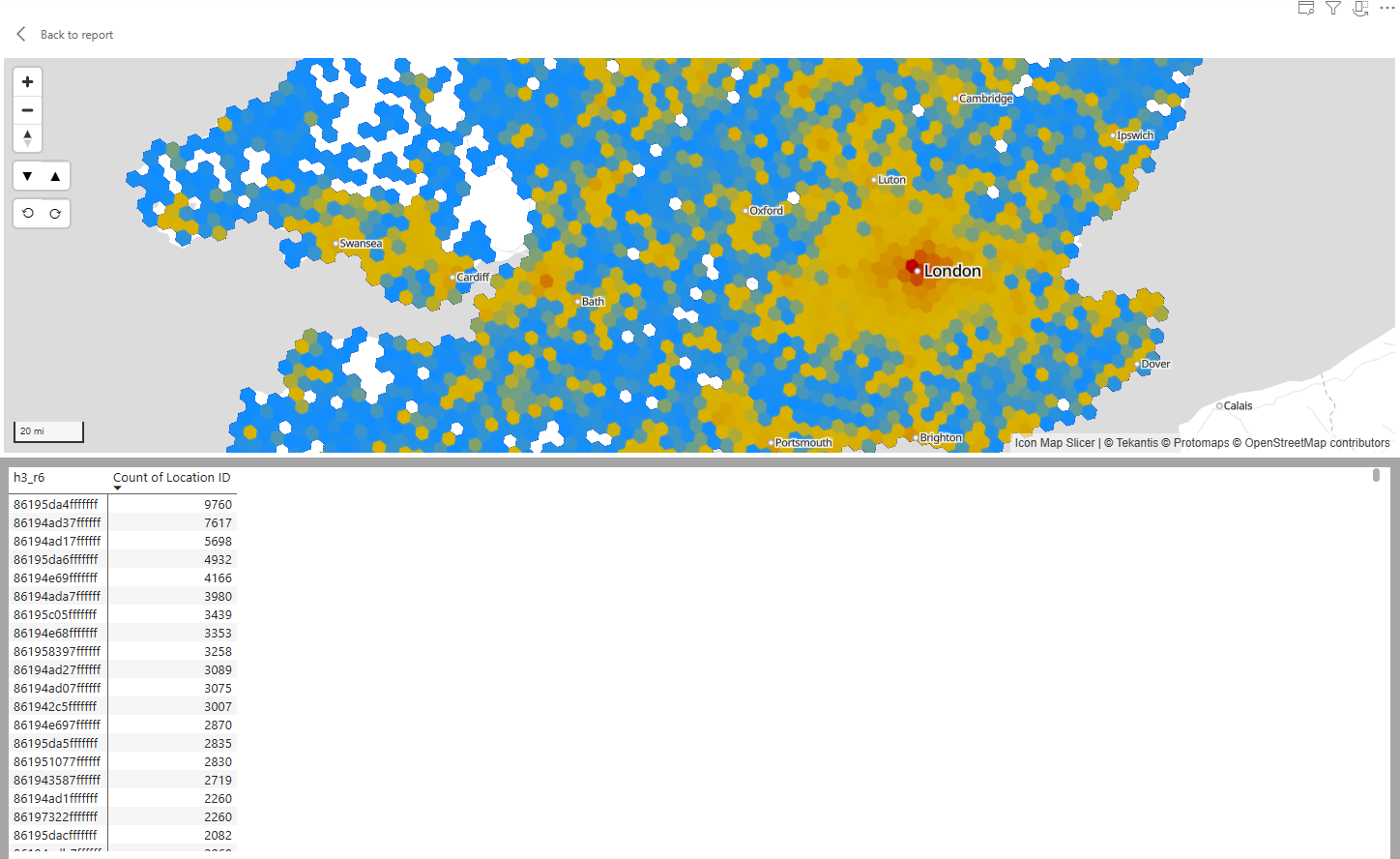
I've created a notebook / python script that will read all the CSV files in a folder containing Longitude and Latitude coordinates and output them to a new folder, with additional columns added for H3 cell IDs at a range of resolutions.
You can download the script from here.
To run it locally:
python add_h3_to_csvs.py --input-dir "d:\sourcedata" --output-dir "d:\outputdata" --lat-col Latitude --lon-col Longitude --res-start 5 --res-end 12 --combine-output --combined-filename "all_data.csv"
The script will read all the CSV files in d:\sourcedata and output a single new file d:\outputdata\all_data.csv with 8 new columns for H3 Cell IDs for resolutions 5 to 12.
You can also run this as a Microsoft Fabric Notebook based on files stored in OneLake.
%pip install h3
import os
import glob
from pathlib import Path
import pandas as pd
import h3 # top-level API
def h3_index(lat: float, lon: float, res: int):
"""
Wrapper to handle both old and new h3-py APIs.
"""
if hasattr(h3, "geo_to_h3"): # older API
return h3.geo_to_h3(lat, lon, res)
elif hasattr(h3, "latlng_to_cell"): # newer API (v4+)
return h3.latlng_to_cell(lat, lon, res)
else:
raise AttributeError(
"Installed h3 package has neither geo_to_h3 nor latlng_to_cell. "
"Please check your h3 version."
)
def add_h3_columns(
df: pd.DataFrame,
lat_col: str,
lon_col: str,
res_start: int,
res_end: int
) -> pd.DataFrame:
"""
Add H3 index columns for resolutions [res_start, res_end] (inclusive)
based on latitude/longitude columns in the DataFrame.
Invalid or out-of-range lat/lon rows will get None in the H3 columns.
"""
if lat_col not in df.columns or lon_col not in df.columns:
raise ValueError(f"Latitude/longitude columns '{lat_col}'/'{lon_col}' not found in dataframe.")
# Coerce to numeric; invalid strings become NaN
lats = pd.to_numeric(df[lat_col], errors="coerce")
lons = pd.to_numeric(df[lon_col], errors="coerce")
# Valid range mask
valid_mask = (
lats.notna()
& lons.notna()
& (lats >= -90) & (lats <= 90)
& (lons >= -180) & (lons <= 180)
)
invalid_count = (~valid_mask).sum()
if invalid_count > 0:
print(f" Warning: {invalid_count} rows have invalid lat/lon and will get empty H3 cells.")
lat_arr = lats.to_numpy()
lon_arr = lons.to_numpy()
valid_arr = valid_mask.to_numpy()
for res in range(res_start, res_end + 1):
col_name = f"h3_r{res}"
result = []
for lat, lon, is_valid in zip(lat_arr, lon_arr, valid_arr):
if not is_valid:
result.append(None)
continue
try:
result.append(h3_index(float(lat), float(lon), res))
except Exception:
# In case H3 still complains for some row, fall back to None
result.append(None)
df[col_name] = result
return df
def process_folder(
input_dir: str,
output_dir: str,
lat_col: str,
lon_col: str,
res_start: int,
res_end: int,
combine_output: bool = False,
combined_filename: str = "combined_with_h3.csv",
pattern: str = "*.csv"
):
"""
Process all CSV files in input_dir, add H3 columns, and write outputs.
- If combine_output = False:
writes one output CSV per input CSV into output_dir
- If combine_output = True:
writes a single combined CSV (all files appended) into output_dir/combined_filename
"""
input_path = Path(input_dir)
output_path = Path(output_dir)
output_path.mkdir(parents=True, exist_ok=True)
files = sorted(glob.glob(str(input_path / pattern)))
if not files:
raise FileNotFoundError(f"No files matching pattern '{pattern}' found in {input_dir}")
print(f"Found {len(files)} files to process.")
combined_path = output_path / combined_filename
header_written = False # for combined mode
for fpath in files:
fname = os.path.basename(fpath)
print(f"Processing {fname}...")
# Read CSV from OneLake
df = pd.read_csv(fpath)
# Add H3 columns
df = add_h3_columns(df, lat_col=lat_col, lon_col=lon_col,
res_start=res_start, res_end=res_end)
if combine_output:
df.to_csv(
combined_path,
mode="a",
index=False,
header=not header_written
)
header_written = True
else:
out_file = output_path / fname
df.to_csv(out_file, index=False)
if combine_output:
print(f"Combined output written to: {combined_path}")
else:
print(f"Per-file outputs written to: {output_path}")
# 🔧 CHANGE THESE TO MATCH YOUR LAKEHOUSE + FOLDER NAMES
lakehouse_name = "YourLakehouseName" # e.g. "GeoDataLH"
input_subfolder = "sourcedata" # folder under Files where your CSVs are
output_subfolder = "outputdata" # folder under Files to write results
input_dir = f"/lakehouse/{lakehouse_name}/Files/{input_subfolder}"
output_dir = f"/lakehouse/{lakehouse_name}/Files/{output_subfolder}"
print("Input dir :", input_dir)
print("Output dir:", output_dir)
# Make sure output folder exists
os.makedirs(output_dir, exist_ok=True)
# Run the processing: Latitude / Longitude columns, resolutions 5–12, single combined file
process_folder(
input_dir=input_dir,
output_dir=output_dir,
lat_col="Latitude", # your actual column names
lon_col="Longitude",
res_start=5,
res_end=12,
combine_output=True, # False = per-file; True = one big CSV
combined_filename="all_data_with_h3.csv",
pattern="*.csv"
)
Here's the resulting Power BI report to explore. It has 3 pages:
- H3 from cell IDs at resolution 9 - 120k cells displayed
- H3 from cell IDs at resolution 6 - 4.4k cells displayed
- H3 at resolution 6 from from 210k longitudes and latitudes
And the .pbix file to download.